MolHuiTu — Molecular HyperGraph
Intelligent Drug–Target Interaction (DTI) Prediction Platform
A next-generation, GPU-accelerated DTI system that fuses hypergraph molecular encoders, protein language models (ProtBert), rigorous explainability, and a clean web UI.
Overview
MolHuiTu (Molecular Intelligence Graph) predicts drug–target interactions from a SMILES (ligand) and a FASTA (protein). It returns a calibrated score and explains the prediction by highlighting key atoms and residues. The web UI includes interactive 3D visualization, batch job management, and downloadable reports.
Feature Highlights
- Hypergraph Molecular Encoder — Captures multi-body patterns (rings, functional groups, H-bonds) beyond pairwise bonds via hyperedges and a masked-autoencoder pretrain; improves modeling of complex chemistry.
- Protein Language Model (ProtBert) — Transformer embeddings of amino-acid sequences (mean/CLS pooling), fused with ligand embeddings for robust DTI scoring.
- End-to-End Inference — Single query and high-throughput batch CSV screening; optional probability calibration for deployment realism.
- Integrated Explainability — Atom-level SHAP and residue-level occlusion with Top-K contributors and a consistency check.
- One-Stop Context — Hooks for PubChem / UniProt / AlphaFold / RCSB PDB to enrich reports and drive 3Dmol.js visualization.
- Practical UX — Clean web UI, job history, CSV export, and report pages; GPU-optimized backend validated on NVIDIA RTX 4090.
Traditional graph vs hypergraph: a simple graph restricts bonds to pairs; MolHuiTu uses hyperedges to connect any number of atoms so functional motifs are represented natively.
Guided Tour
Home → Entry Points
A minimal home screen routes to single prediction, batch submission, and history.
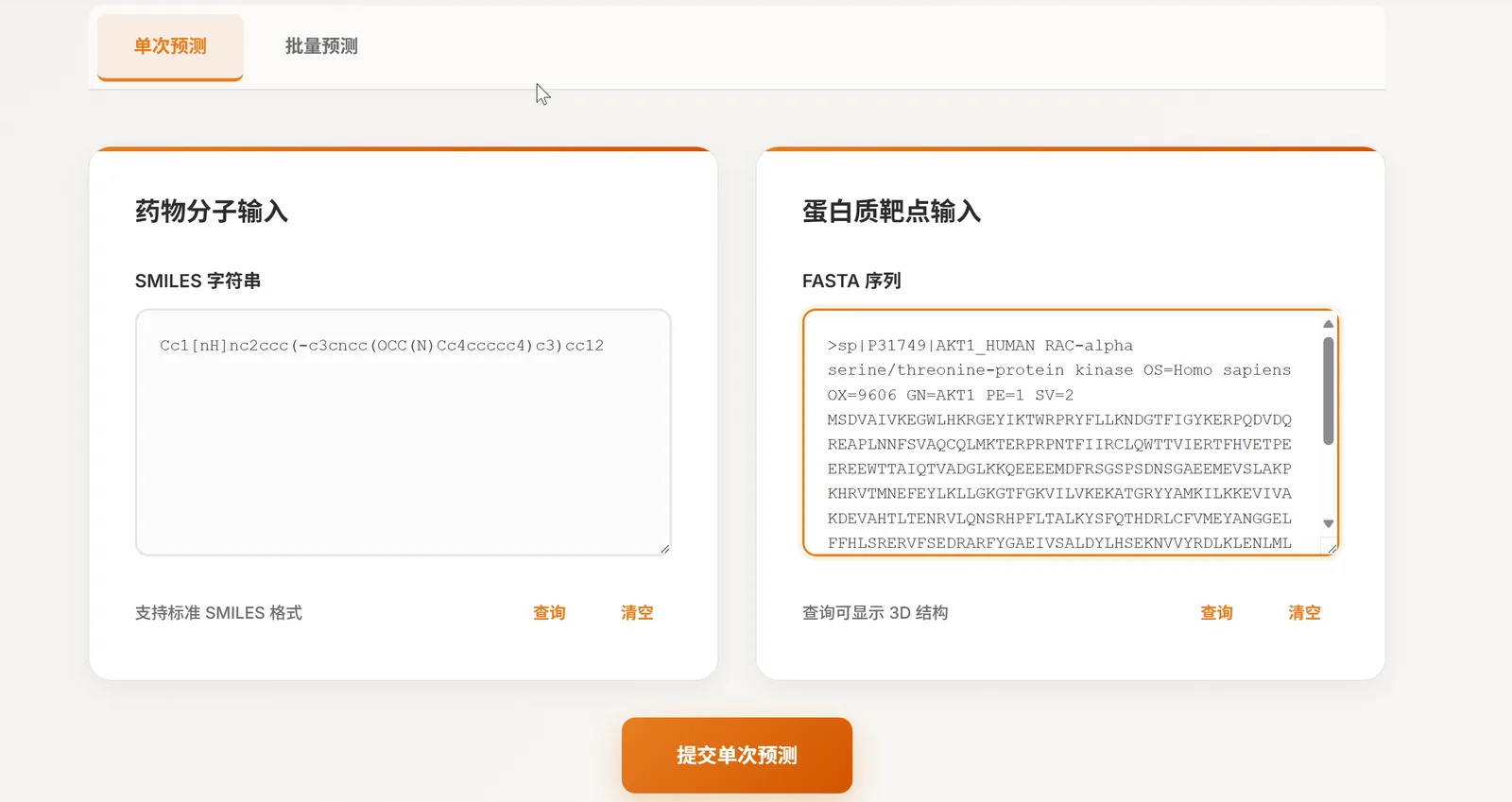
Single Prediction → Fill & Submit
Provide SMILES and a FASTA (or UniProt ID). Toggle explainability if needed.
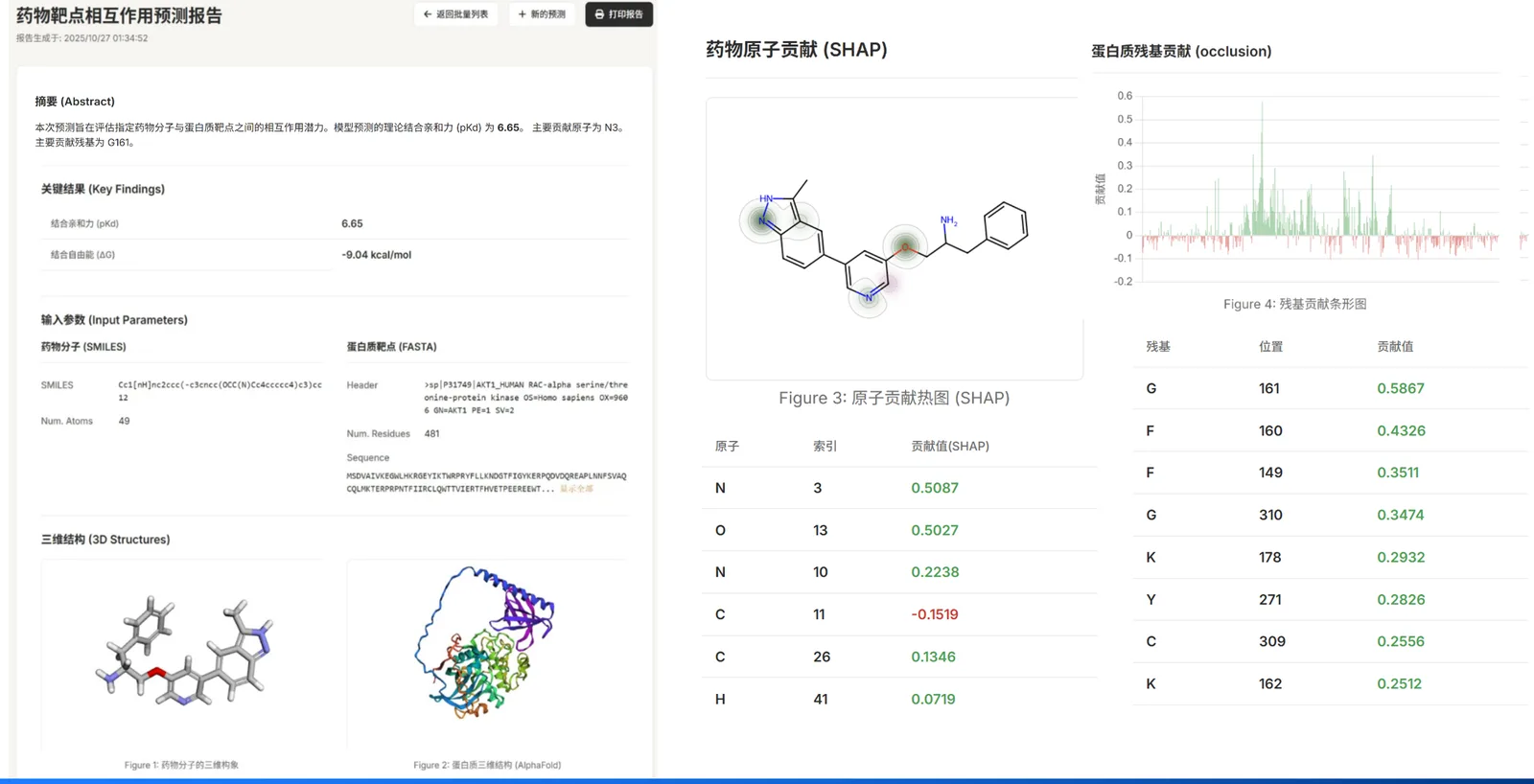
Reports → Scores & Rationale
Per-sample reports summarize inputs, prediction scores, and visuals.
Architecture
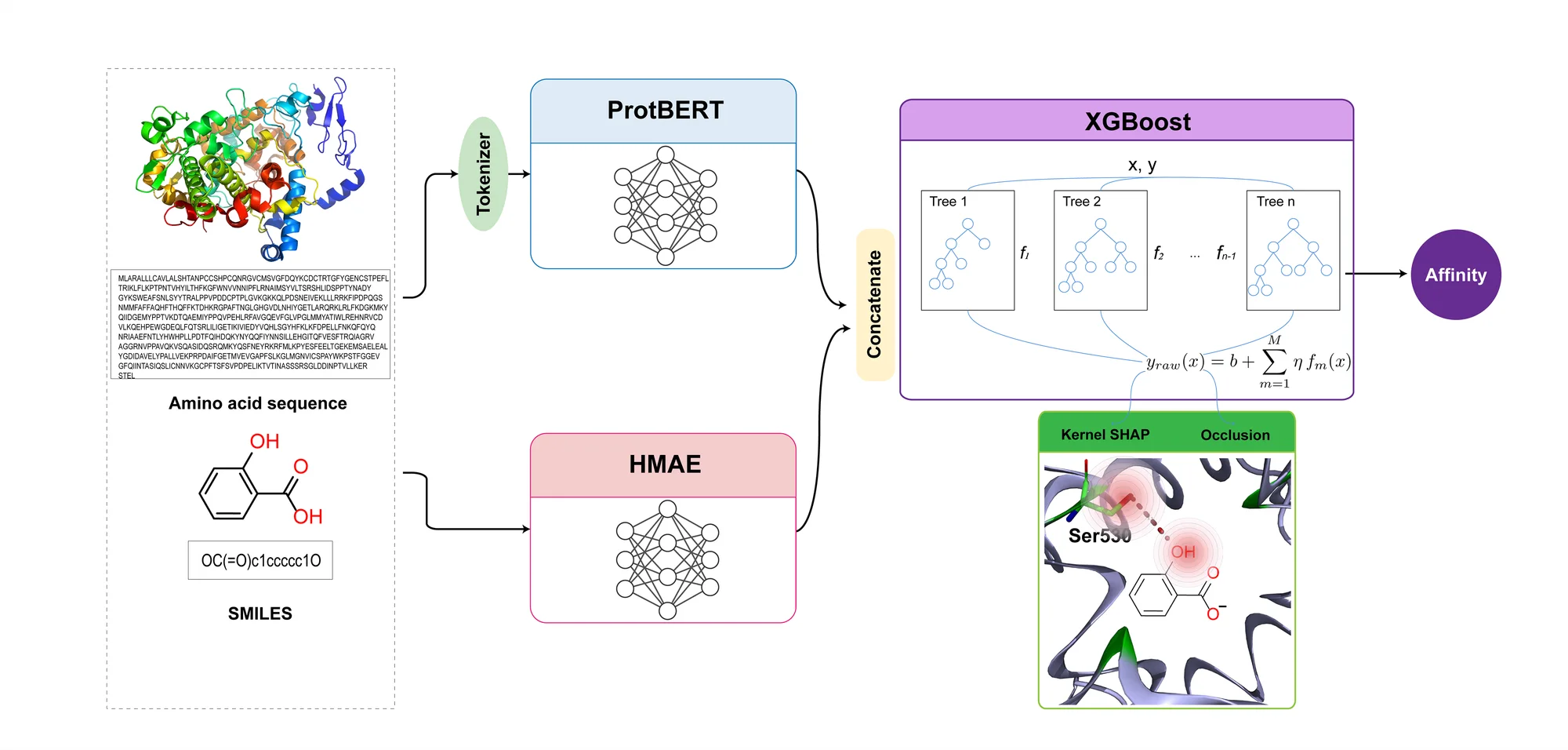
- Drug Encoder — HyperGraph-MAE: Represent molecules as hypergraphs (nodes = atoms; hyperedges = rings/groups/relations). Pretrain with degree-aware masking and reconstruction; aggregate via multi-head attention → fixed-size ligand embedding.
- Protein Encoder — ProtBert: Transformer embeddings from ProtBert (HuggingFace); mean/CLS pooling configurable → protein embedding.
- Fusion & Prediction — XGBoost Head: Concatenate (or bilinear fuse) ligand/protein embeddings → XGBoost for classification (probability) or regression (affinity). Optional Platt / Isotonic calibration improves reliability.
Backend stack: PyTorch (+ CUDA), PyTorch Geometric, RDKit, XGBoost, SHAP, 3Dmol.js (frontend).